>> Research
| Publications
| Google Scholar
| Pubmed
| ORCID
-> PREPRINT New preprint by my first PhD student Abrar Bhat (at NCBS) on the sorting of membrane proteins by local actomyosin contractility.
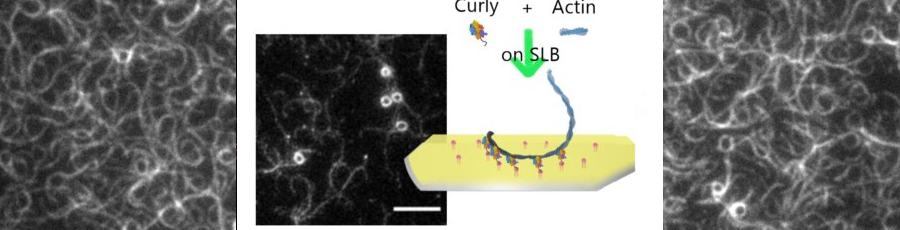
-> NEW ARTICLE New paper from our lab in collaboration with Richard Morris at UNSW and EMBL Australia!
 Darius Köster (group leader, contact, @ActinDomain, LinkedIn) |CV
Darius Köster (group leader, contact, @ActinDomain, LinkedIn) |CV
How do cells cope with their environment? Deciphering mechanical processes at the cell surface leading to signalling events and the adaptation of cells to changes in the environment.
My research interests lie in the understanding the molecular and physical principles that govern processes at the plasma membrane of cells. Particularly, by which mechanisms the force generating machinery of the cell cortex,
structural filaments and motor proteins, govern and regulate the mechanical properties of the cell membrane and dynamics of cell membrane components, and vice versa, how membrane organisation and signalling events feed-back
to the regulation of the cortex machinery. These mechanisms, which in turn regulate cell motility and cell-cell interactions, underlie important, poorly-understood human diseases that constitute global health problems.
The lab employs novel assays based on reconstituted membrane systems in combination with measurements on live cells using state of the art fluorescence microscopy and mechanical manipulation.
 John James (funded by EPSRC, LinkedIn)
John James (funded by EPSRC, LinkedIn)
I am a post-doctoral researcher with extended experience in cell biology, genetic engineering and microscopy, supplemented by basic knowledge of biochemistry and computational data analysis. I utilise this skill-set to study the interplay between actin-generated forces and the maturation of cell adhesions.
During my PhD at the lab of Alexis Gautreau (Ecole Polytechnique, France), I showed that vinculin, a mechano-sensor present at focal adhesions and adherens junctions, suppresses the generation of branched actin networks. This signal impacts several cell behaviours ranging from proliferation to migration to attachment, potentially establishing a novel tumor-suppressor pathway. I now join the Koester Lab with the hope of understanding how forces generated by these branched actin networks can in turn influence the maturation and stability of cell adhesions. Uncovering such an inverse regulation could complete a feedback loop essential for the sustenance of cell adhesion and thus the prevention of cancer metastasis.
---PhD Candidates---
 Yugesh Ramdhun (MIBTP)
Yugesh Ramdhun (MIBTP)
My interest in mechanobiology began with deciphering how cells adapt to mechanical changes in their micro-environment and cell mechanoreciprocity. I studied the movement of extracellular vesicles in strained microenvironments, uncovering their role in ECM remodelling through degradation during my master’s at IIT Madras in the Cell Mechanics Lab. For my PhD, I aim to investigate the link between cell adhesion to collagen in connective tissue and the development of hypermobile Ehlers-Danlos Syndrome (hEDS). Specifically, how integrin-mediated adhesion to ECM components modulates the mechanical properties of fibroblasts, and how these cellular mechanics reciprocally influence adhesion. My research will also focus on fibroblast contractility and traction forces, comparing wild-type fibroblasts with cell lines derived from patients with hypermobile and classical EDS to gain mechanistic insights into the disease. Additionally, I will use atomic force microscopy to investigate how fibroblasts remodel the surrounding ECM and alter its mechanical properties.
 Lucie Winn (MRC-DTP)
Lucie Winn (MRC-DTP)
My PhD project focusses on cell-cell adhesion, particularly via E-cadherin. Cadherins can recruit adapter proteins which link the cell-cell adhesion site to the actomyosin cytoskeleton of cells. This is a mechanosensitive process, and factors such as E-cadherin mobility impact regulation of the cytoskeleton. However, the key mechanisms and cascades of such mechanosensitive signalling remain unknown. During my PhD I hope to gain a better understanding of this process, and delve into questions such as, is mobility-mediated regulation local (at contact site only) or global (translated over the entire cell surface)? How are different proteins activated to organise the cytoskeleton? And how do other mechanical constraints affect E-cadherin mechano-signalling? We hope a better understanding of this fundamental biological process will provide important insights into several other related biological process, such as tissue development, wound healing and cancer metastasis.
 Badeer Hassan Ummad (Chancellor International Scholarship, LinkedIn, @BadeerHassan)
Badeer Hassan Ummad (Chancellor International Scholarship, LinkedIn, @BadeerHassan)
My research interest lies in integrating quantitative techniques in biology to understand complex cellular phenomena. I started exploring biology during my master at IISER Pune by studying cytoskeletal organisation inside cellular geometry using computational modelling approaches. These experiences got me fascinated with the applications of quantitative methods to study biology. At Koester lab, I will be looking into the interaction of actin cytoskeleton with cellular membrane. This research will help to learn how the cytoskeletal network organisation determines cell shape and how biochemical and mechanical signals control this process.
----------------<3 Alumni <3----------------
Sabeeha Malek (completed in 2025, MIBTP)
PhD thesis - The role of cell adhesion and cytoskeleton dynamics in the pathogenesis of Ehlers-Danlos syndrome)
Continued with a postdoc in our lab till Dec. 2025.
Lea Denker (completed in 2025, MIBTP)
PhD thesis - Discerning the molecular mechanisms surrounding membrane rearrangement into double membrane vesicles by SARS-CoV-2 non-structural proteins 3 and 4)
Continued as Associate Medical writer at Healthcare Consultancy Group
Paulina Pokorska (completed in 2025, ARAP)
PhD thesis - Using APEX2 to understand septate junction maturation in Drosophila embryo)
Continued with a postdoc in the lab of Prof. Li, MBI Singapore.
Sayantika Ghosh (completed in 2025, Chancellor International Scholarship)
Thesis title: Real-time Analysis of E-Cadherin mediated Junction Formation in Cancer Cells Using a Minimal Reconstituted System.
Continued with a postdoc in the lab of Laura Machesky, University of Cambridge.
Post-docs
- Sedigheh (Mobi) Ghanbarzadeh Nodehi, 2022-25, now in the laboratory of Prof. Mayor, WMS
- Zoe Shofield, 2022, now at University of Birmingham
- Scott Clarke, 2020-21, now at WMG
---Master Students---
- Peter Mottram-Epson (Medical Sciences MRes, 2024-25)
Thesis title: Investigating Adherens Junction formation and changes in cell shape within cells on supported lipid bilayers under dynamic confinement.
Continued as a PhD student in the Lab Adhesion Inflammation) at Aix-Marseille University, France.
project students
- Sarah Happ (visiting from University of Geneva)
- Alina Makhmudova (visiting from University of Geneva)
- Deepak Ishwara Shivaprakash (visiting from PES University Bangalore)
----------------------------------------------